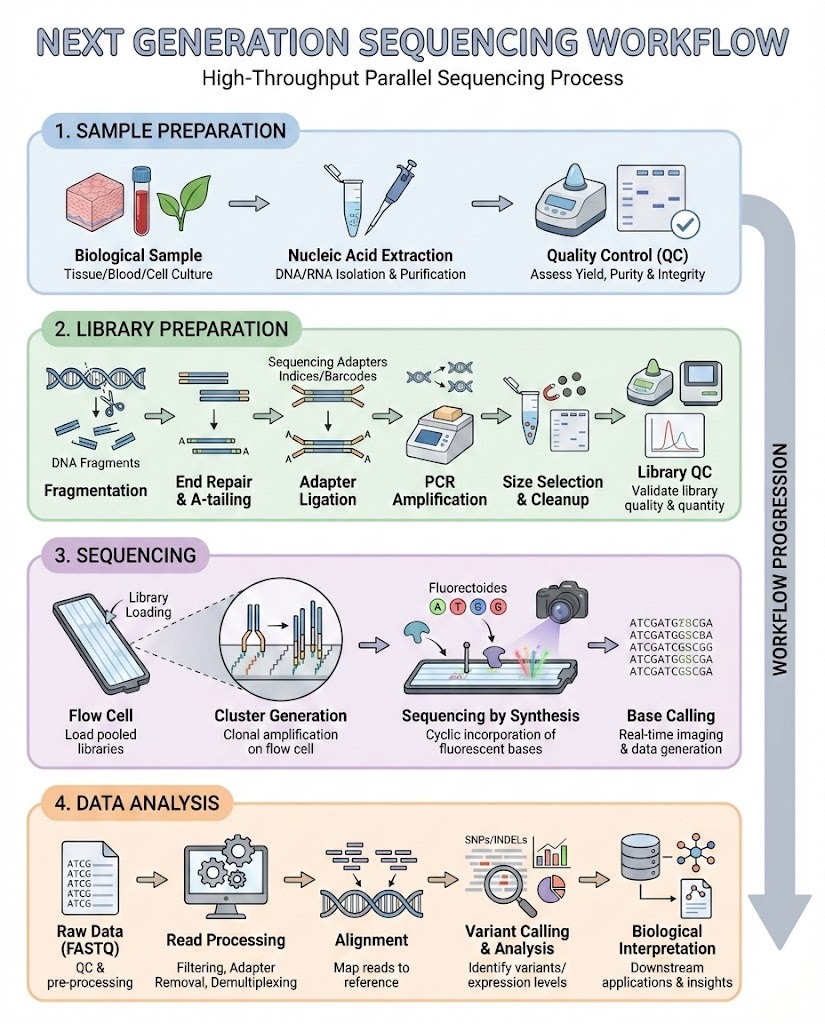
Next generation sequencing has transformed genomics by enabling rapid, high-throughput analysis of DNA and RNA. At the center of this technology is the Next Generation Sequencing Workflow, a structured process that converts biological samples into actionable genomic data.
Understanding this workflow is essential for researchers, clinical laboratories, and biotech professionals who want reliable results, regulatory compliance, and efficient operations. This guide walks through each stage of the workflow, highlights best practices, and explains how optimization improves data quality and downstream analysis.
What Is the Next Generation Sequencing Workflow?
The Next Generation Sequencing Workflow refers to the complete sequence of steps required to generate and analyze sequencing data. It begins with sample preparation and ends with data interpretation.
Each step is tightly connected. Errors early in the process can compromise the final dataset, while optimized workflows improve:
-
Accuracy and reproducibility
-
Turnaround time
-
Cost efficiency
-
Data reliability for clinical or research use
A well-designed workflow also ensures compatibility with sequencing platforms and bioinformatics pipelines.
Core Stages of the Next Generation Sequencing Workflow
While protocols vary by application, most workflows follow the same fundamental structure.
1. Sample Collection and Quality Assessment
Everything begins with high-quality biological material. Common sample types include:
-
Blood and tissue
-
Saliva or buccal swabs
-
Cultured cells
-
Microbial or environmental samples
Key quality checks include:
-
DNA or RNA concentration
-
Purity ratios
-
Fragment integrity
Poor sample quality leads to biased libraries and unreliable sequencing output.
2. Nucleic Acid Extraction
Extraction isolates DNA or RNA from the sample while removing proteins, lipids, and inhibitors. Laboratories typically use:
-
Column-based purification kits
-
Magnetic bead systems
-
Automated extraction platforms
Consistency at this stage is critical for scalable and repeatable sequencing projects.
3. Library Preparation
Library preparation converts extracted nucleic acids into fragments compatible with sequencing instruments. This step usually involves:
-
Fragmentation
-
End repair and adapter ligation
-
PCR amplification
-
Size selection
Library preparation is often the most technically demanding part of the Next Generation Sequencing Workflow because it directly influences:
-
Coverage uniformity
-
Read length
-
Error rates
Automation is increasingly used to minimize variability and contamination.
4. Sequencing
Prepared libraries are loaded onto a sequencing platform such as Illumina, Ion Torrent, or Oxford Nanopore. During sequencing:
-
DNA fragments are read base by base
-
Millions to billions of reads are generated
-
Raw data is stored as FASTQ files
Platform choice depends on:
-
Required read length
-
Throughput
-
Budget
-
Application type such as whole genome, targeted panels, or RNA sequencing
5. Data Processing and Quality Control
Raw sequencing data must be processed before analysis. This stage includes:
-
Base calling
-
Read trimming and filtering
-
Removal of adapters
-
Quality score assessment
Common quality metrics include:
-
Q30 scores
-
Read depth
-
Duplication rates
-
GC bias
These checks ensure that downstream analysis is based on reliable data.
6. Bioinformatics Analysis
This step transforms raw reads into meaningful biological insights. Depending on the project, analysis may involve:
-
Alignment to a reference genome
-
Variant calling
-
Gene expression quantification
-
Metagenomic classification
Specialized software and pipelines are used to automate processing while maintaining reproducibility.
7. Interpretation and Reporting
Final results are interpreted by researchers or clinicians. In clinical settings, this may include:
-
Identifying pathogenic variants
-
Assessing treatment options
-
Generating regulatory-compliant reports
Clear documentation and validation are critical at this stage.
Why Workflow Optimization Matters
An optimized Next Generation Sequencing Workflow reduces technical failures and improves data confidence. Key benefits include:
-
Lower repeat rates due to failed libraries
-
Faster turnaround times
-
Reduced reagent waste
-
Better reproducibility between runs
Optimization strategies often focus on:
-
Standard operating procedures
-
Automation
-
Quality control checkpoints
-
Workflow documentation
These improvements are especially valuable for high-throughput laboratories and clinical diagnostics.
Common Applications of the Workflow
The flexibility of next generation sequencing enables use across many disciplines.
Popular applications include:
-
Cancer genomics and tumor profiling
-
Rare disease diagnosis
-
Infectious disease surveillance
-
Agricultural genomics
-
Microbiome research
-
Drug development and pharmacogenomics
Each application may adjust certain workflow steps, but the core structure remains the same.
Choosing Tools and Platforms for Your Workflow
Selecting the right tools impacts performance and cost.
Important considerations include:
-
Compatibility between library kits and sequencers
-
Availability of technical support
-
Regulatory compliance for clinical use
-
Integration with laboratory information systems
-
Scalability for future growth
Many laboratories evaluate complete workflow solutions from a single vendor to reduce compatibility issues.
Challenges in Implementing NGS Workflows
Despite its advantages, the workflow presents several challenges:
-
High initial equipment costs
-
Complex data management
-
Requirement for skilled personnel
-
Ongoing validation and quality assurance
Addressing these challenges requires careful planning and continuous training.
Best Practices for Reliable Sequencing Results
To maintain high-quality outcomes:
-
Use standardized protocols
-
Implement routine quality control
-
Track reagent lot numbers
-
Validate new methods before clinical use
-
Maintain detailed documentation
These practices support long-term reliability and regulatory compliance.
Conclusion
The Next Generation Sequencing Workflow is the foundation of modern genomic analysis. From sample collection to data interpretation, each stage contributes to the accuracy and usefulness of sequencing results.
Organizations that invest in workflow optimization benefit from higher data quality, faster project completion, and greater confidence in their findings. Whether used for clinical diagnostics, research, or biotechnology development, a structured and well-managed workflow ensures that next generation sequencing delivers its full potential.
Frequently Asked Questions
What are the main steps in a next generation sequencing workflow?
The main steps include sample collection, nucleic acid extraction, library preparation, sequencing, data processing, bioinformatics analysis, and result interpretation. Each step builds on the previous one to produce reliable genomic data.
How long does a typical NGS workflow take?
Depending on the application and platform, the workflow can take anywhere from one to several days. Library preparation and sequencing time are usually the most time-consuming stages.
What factors affect the quality of an NGS workflow?
Sample quality, library preparation accuracy, sequencing platform performance, and bioinformatics pipeline design all play major roles in determining overall data quality.
Is the next generation sequencing workflow suitable for clinical diagnostics?
Yes. Many clinical laboratories use validated NGS workflows for cancer profiling, genetic disease testing, and infectious disease detection, provided they meet regulatory and quality standards.