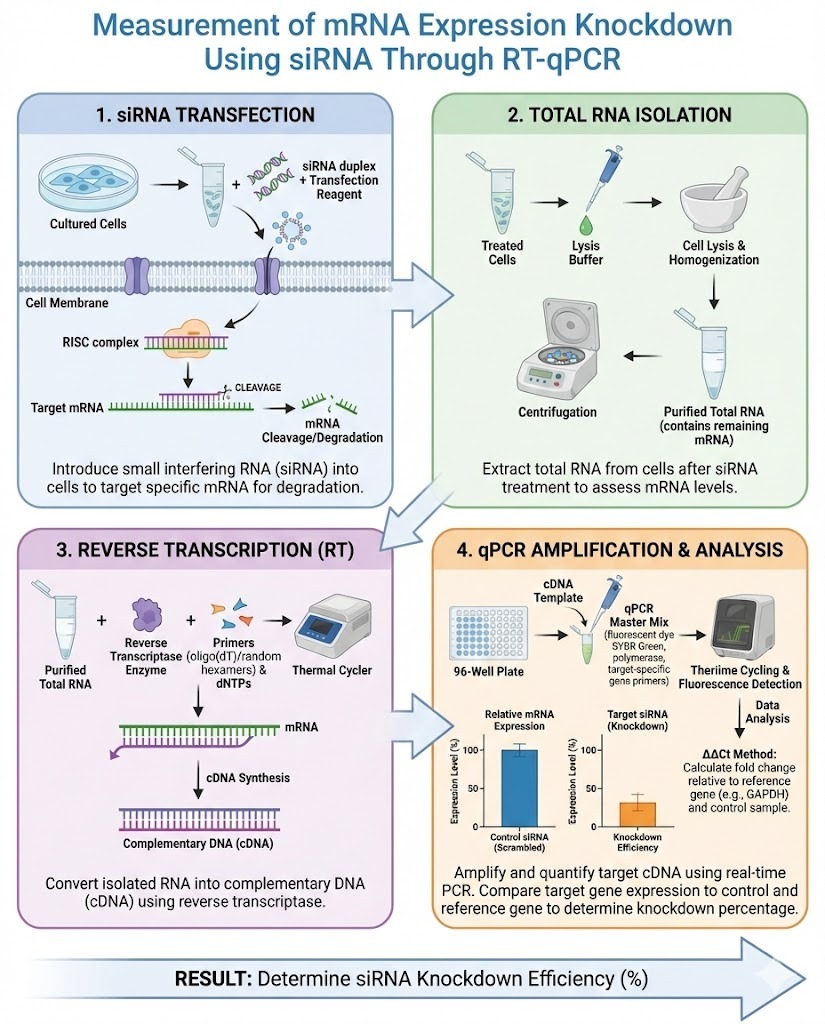
Gene silencing using small interfering RNA has become a standard technique in molecular biology for studying gene function and validating therapeutic targets. However, the success of any siRNA experiment depends on accurately determining how much target gene expression has been reduced. The Measurement of mRNA Expression Knockdown Using siRNA Through RT-qPCR is widely regarded as the most sensitive and quantitative method for this purpose.
This article explains the scientific rationale, workflow, technical considerations, and best practices for assessing siRNA-mediated knockdown using reverse transcription quantitative PCR. It is designed for researchers seeking reliable, reproducible, and publication-quality results.
Why Measure mRNA Knockdown After siRNA Treatment
siRNA works by promoting the degradation of specific mRNA molecules, thereby reducing protein production. Without proper measurement, it is impossible to know whether:
-
The siRNA successfully entered the cells
-
The target transcript was efficiently degraded
-
Off-target effects occurred
-
Experimental conclusions are valid
Protein-level assays such as western blotting are useful, but mRNA-level analysis provides earlier and more direct evidence of gene silencing. This is why the measurement of mRNA expression knockdown using siRNA through RT-qPCR has become a gold standard in gene expression studies.
Overview of the RT-qPCR-Based Knockdown Assessment Workflow
Although protocols vary between laboratories, the core workflow is consistent.
Major stages include:
-
Cell treatment with siRNA
-
RNA isolation
-
Reverse transcription
-
Quantitative PCR amplification
-
Data normalization and analysis
Each stage must be carefully controlled to ensure accurate quantification of gene expression changes.
Step 1: siRNA Transfection and Experimental Design
Effective measurement begins with thoughtful experimental planning.
Key considerations include:
-
Selection of validated siRNA sequences
-
Use of appropriate negative controls such as scrambled siRNA
-
Inclusion of untreated controls
-
Optimization of siRNA concentration
-
Timing of sample collection post-transfection
Most experiments measure mRNA levels 24 to 72 hours after siRNA delivery, depending on cell type and gene turnover rate.
Biological replicates are essential for statistical reliability.
Step 2: RNA Extraction and Quality Assessment
After treatment, total RNA is extracted from cells using column-based or reagent-based purification systems.
High-quality RNA is critical for reliable RT-qPCR results.
Important quality checks include:
-
RNA concentration measurement
-
Purity ratios such as A260/A280
-
Assessment of RNA integrity
Degraded RNA leads to underestimation of transcript levels and increased variability.
Step 3: Reverse Transcription to cDNA
Extracted RNA is converted into complementary DNA using reverse transcriptase enzymes.
Best practices include:
-
Using equal RNA input amounts for all samples
-
Including no-reverse-transcriptase controls
-
Selecting appropriate primers such as random hexamers or oligo-dT
The consistency of this step directly affects quantification accuracy in the measurement of mRNA expression knockdown using siRNA through RT-qPCR.
Step 4: Quantitative PCR Amplification
Quantitative PCR amplifies the cDNA corresponding to the target gene and reference genes.
Important technical elements include:
-
Primer specificity and efficiency
-
Reaction chemistry such as SYBR Green or probe-based systems
-
Thermal cycling conditions
-
Instrument calibration
Target gene expression is measured relative to internal reference genes, commonly referred to as housekeeping genes.
Step 5: Data Normalization and Analysis
Raw Ct values must be processed to calculate relative expression levels.
Common analysis steps:
-
Normalize target gene Ct values to reference genes
-
Compare treated and control samples
-
Calculate fold change using standard mathematical models
-
Perform statistical testing
This analysis yields the percentage or fold reduction in mRNA levels, which defines the knockdown efficiency.
Choosing Reference Genes for Accurate Quantification
Reference gene selection is one of the most important factors in RT-qPCR accuracy.
Ideal reference genes should:
-
Be stably expressed across conditions
-
Not be affected by siRNA treatment
-
Show minimal variation between replicates
Using more than one reference gene further improves reliability.
Common Sources of Error and How to Avoid Them
Even experienced laboratories encounter challenges when performing the measurement of mRNA expression knockdown using siRNA through RT-qPCR.
Frequent issues include:
-
Incomplete transfection efficiency
-
RNA degradation
-
Primer-dimer formation
-
Variable reference gene expression
-
Pipetting errors
Preventive strategies:
-
Optimize transfection conditions for each cell line
-
Use RNase-free reagents and consumables
-
Validate primer specificity
-
Run technical replicates
-
Implement strict laboratory protocols
Interpreting Knockdown Results
Successful knockdown typically shows:
-
A statistically significant reduction in target mRNA
-
Dose-dependent effects when multiple siRNA concentrations are tested
-
Consistency across biological replicates
Partial knockdown is common and often sufficient for functional studies. However, extremely low efficiency may indicate poor siRNA design or delivery issues.
Applications of RT-qPCR Knockdown Measurement
This approach is used across many research areas.
Examples include:
-
Functional genomics
-
Cancer biology
-
Drug target validation
-
Pathway analysis
-
RNA interference screening
Pharmaceutical and biotechnology companies also rely on this method during early-stage therapeutic development.
Advantages of Using RT-qPCR for Knockdown Analysis
RT-qPCR remains the preferred method for several reasons:
-
High sensitivity
-
Wide dynamic range
-
Quantitative output
-
Relatively low cost
-
Compatibility with many sample types
These advantages make it suitable for both small academic projects and large-scale screening studies.
Limitations to Consider
While powerful, this approach has limitations.
-
Does not directly measure protein levels
-
Sensitive to experimental variability
-
Requires careful normalization
-
Cannot detect functional protein activity
For comprehensive studies, RT-qPCR data are often combined with protein assays or phenotypic analyses.
Best Practices Summary
To achieve reliable results:
-
Use validated siRNA reagents
-
Optimize transfection efficiency
-
Ensure high-quality RNA extraction
-
Select stable reference genes
-
Include appropriate controls
-
Perform replicate experiments
-
Document all procedures thoroughly
These steps support consistent measurement of mRNA expression knockdown using siRNA through RT-qPCR across experiments and laboratories.
Conclusion
The Measurement of mRNA Expression Knockdown Using siRNA Through RT-qPCR is a cornerstone technique in gene function studies and therapeutic research. It provides a sensitive, quantitative, and reproducible way to evaluate the effectiveness of gene silencing strategies.
When implemented with careful experimental design, rigorous quality control, and proper data analysis, this method delivers reliable insights into gene regulation and cellular response. For researchers seeking to validate siRNA performance or explore molecular pathways, RT-qPCR remains an indispensable analytical tool.
Frequently Asked Questions
What is the purpose of measuring mRNA knockdown using RT-qPCR after siRNA treatment?
The goal is to quantitatively determine how effectively the siRNA reduces the target gene transcript, confirming successful gene silencing at the molecular level.
How accurate is the measurement of mRNA expression knockdown using siRNA through RT-qPCR?
When properly controlled, RT-qPCR provides highly accurate and reproducible quantification across a wide range of expression levels.
How soon after siRNA transfection should mRNA knockdown be measured?
Most experiments assess knockdown between 24 and 72 hours post-transfection, depending on the cell type and gene stability.
Can RT-qPCR results alone confirm functional gene silencing?
RT-qPCR confirms transcript reduction but does not measure protein levels or biological activity. Protein assays and functional studies are often used to complement mRNA data.